Protein Affinity Chromatography
To purify a protein with the help of their affinity to another molecule, the interacting partner (ligand), e.g., a protein, a small molecule, or a metal, is immobilized on the stationary phase of the chromatography matrix. The stationary phase mostly consists of agarose or synthetic polymers and is packed into a column in the form of beads. The beads are surrounded by a liquid, called the mobile phase. When the target protein-containing sample is applied, it enters the mobile phase and runs through the beads of the stationary phase. Meanwhile, the target protein can bind to the ligand, whereas other molecules remain in the mobile phase and can be removed by washing. For elution of the target protein, the interaction to the ligand is resolved by changing the buffer conditions, for example, the pH value, or by adding a specific competitor that supplants the target protein from the ligand resulting in dissociation of the target, while the ligand remains immobilized on the stationary phase.
Such a direct affinity purification strategy is most commonly used for antibodies based on antigen-antibody interactions. One example is protein A (ligand), which can be utilized to purify immunoglobulin G antibodies (target protein). But how can a protein be purified if an interaction partner is not known so far? In this case, the affinity of known interactions can be utilized for the indirect capture of the target protein. For this purpose, the target protein is tagged with a short peptide of one of the known interactors, which can bind to the other interaction partner immobilized on the stationary phase. The short peptide of the known interactor is called affinity tag and can be, for example, the His-tag, GST-tag, Strep-tag®II, or Twin-Strep-tag®. They can bind to ligands such as metal ions, glutathione, Strep-Tactin® or Strep-Tactin®XT.
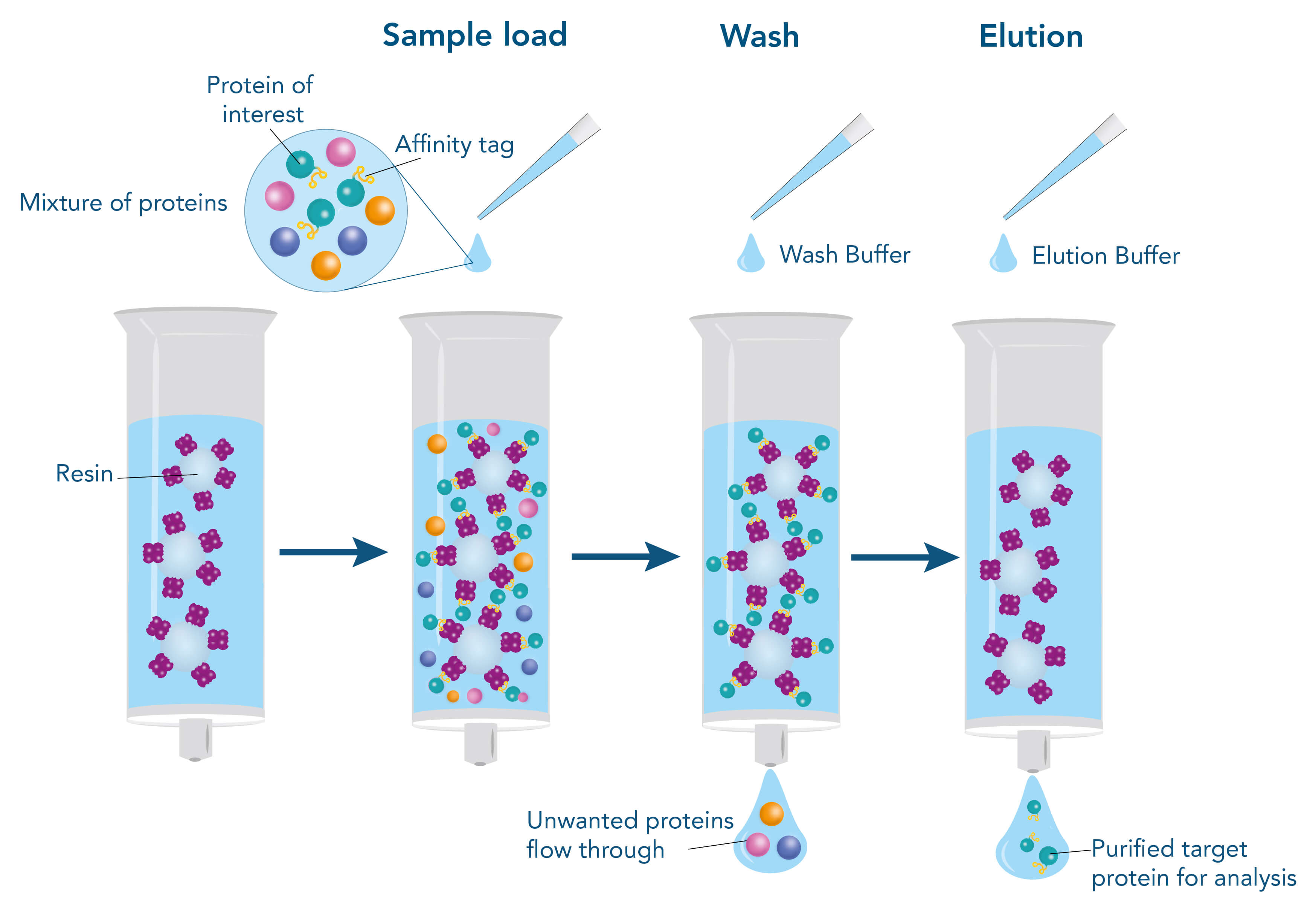
Principle of affinity-based protein purification
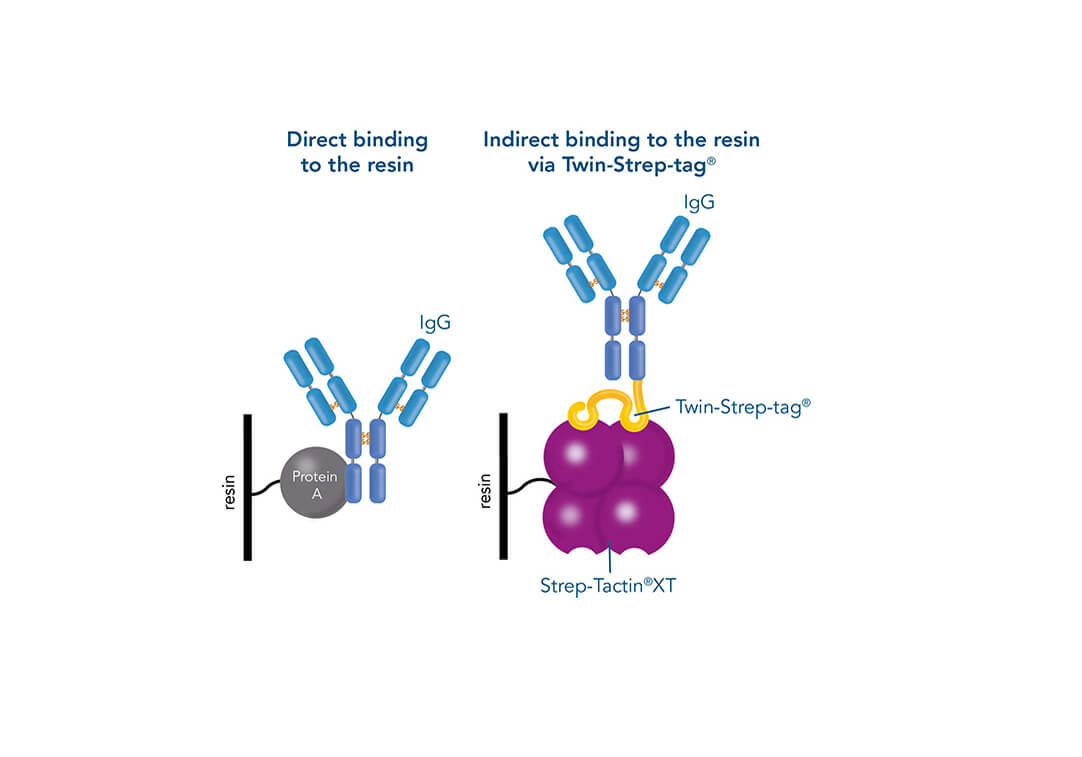
Comparison of direct and indirect affinity purification strategies
Due to the highly specific selection and the resulting purity of target proteins, affinity-based systems have become the flagship method of purification. However, some of the widely used affinity tags, such as the His-tag, exhibit several drawbacks and limitations, since they can increase the risk of distorting the natural conformation of the target protein or necessitate stringent elution and wash conditions affecting the yield of the target protein. Additionally, many tags are not compatible with varying buffer conditions and must be removed to not impair downstream processing.
The widely used Strep-tag® technology, consisting of two engineered streptavidin variants, Strep-Tactin® and Strep-Tactin®XT, and two affinity tags, Strep-tag®II and Twin-Strep-tag®, is not limited in the use of buffers and its high specificity leads to obtaining highly pure proteins.
Therefore, IBA provides several resins, either coupled with Strep-Tactin® or Strep-Tactin®XT, which are all applicable for Strep-tag®II and Twin-Strep-tag® fusion protein purification. The purification cycle varies between both streptavidin variants, but both resin types serve the same goal: simple, fast, and variable protein purification procedures for highly pure proteins.
The following application notes provide comparison of Strep-Tactin® and Strep-Tactin®XT purification systems: for purification of Latex Clearing Protein , for further proteins (1, 2).
The special differences to other systems, especially between the His-tag system and Strep-tag® technology, are presented in a comprehensive comparison. It summarizes the differences as well as recommends one of the systems depending on the properties of the target protein, expression host, and purification conditions.
The differences between systems in protein purification from Expi supernatants is demonstrated in this application note.
IBA’s unique Strep-tag® technology is a commonly used tool for the affinity purification of recombinant proteins and is based on one of the strongest non-covalent interactions in nature, which is the interaction of biotin to streptavidin. The system includes two affinity tags: Strep-tag®II and Twin-Strep-tag® (the tandem version of the Strep-tag®II). These peptide sequences exhibit intrinsic affinity towards two specifically engineered streptavidin variants — Strep-Tactin® and Strep-Tactin®XT.
The Strep-tag® technology stands for the highest protein purities under physiological conditions and can be used in a variety of different applications: in a field of purification, immobilization of proteins for assay development, and for protein interaction studies.

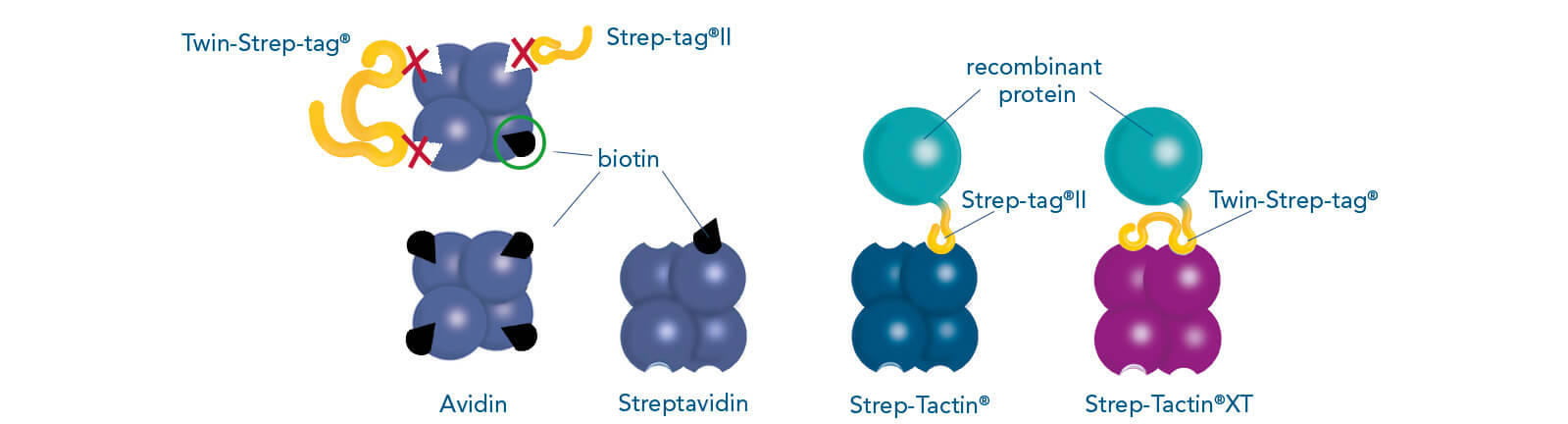
Binding patterns of Strep-tag®II, Twin-Strep-tag® and biotin

